Example 2: reptile-mammals
![[Figure1.4.1 (cartoon of vertebrate jaws)]](http://www.talkorigins.org/faqs/comdesc/images/jaws2.gif) |
Figure 1.4.1. The jaws of three vertebrates—mammal, therapsid, and pelycosaur. A side view of three idealized skulls of mammals, therapsids (mammal-like reptiles), and pelycosaurs (early reptiles). The figure shows the differences between mammal and reptilian jaws and ear-bone structures. The jaw joint is shown as a large black dot, the quadrate (mammalian anvil or incus) is in turquoise, the articular (mammalian hammer or malleus) is in yellow, and the angular (mammalian tympanic annulus) is in pink. Note how, in the reptile, the jaw joint is formed between the blue quadrate and the yellow articular (with the pink angular close by), and how, in the mammal, the jaw joint is formed between the squamosal above and the dentary below. In the reptile, the squamosal is just above and contacting the quadrate. Advanced therapsids have two jaw joints: a reptile-like joint and a mammal-like joint (Figure based on Kardong 2002, pp. 275, reproduced with permission from the publisher, Copyright © 2002 McGraw-Hill) |
We also have an exquisitely complete series of fossils for the reptile-mammal intermediates, ranging from the pelycosauria, therapsida, cynodonta, up to primitive mammalia (Carroll 1988, pp. 392-396; Futuyma 1998, pp. 146-151; Gould 1990; Kardong 2002, pp. 255-275). As mentioned above, the standard phylogenetic tree indicates that mammals gradually evolved from a reptile-like ancestor, and that transitional species must have existed which were morphologically intermediate between reptiles and mammals—even though none are found living today. However, there are significant morphological differences between modern reptiles and modern mammals. Bones, of course, are what fossilize most readily, and that is where we look for transitional species from the past. Osteologically, two major striking differences exist between reptiles and mammals: (1) reptiles have at least four bones in the lower jaw (e.g. the dentary, articular, angular, surangular, and coronoid), while mammals have only one (the dentary), and (2) reptiles have only one middle ear bone (the stapes), while mammals have three (the hammer, anvil, and stapes) (see Figure 1.4.1).
Early in the 20th century, developmental biologists discovered something that further complicates the picture. In the reptilian fetus, two developing bones from the head eventually form two bones in the reptilian lower jaw, the quadrate and the articular (see the Pelycosaur in Figure 1.4.1). Surprisingly, the corresponding developing bones in the mammalian fetus eventually form the anvil and hammer of the unique mammalian middle ear (also known more formally as the incus and malleus, respectively; see Figure 1.4.2) (Gilbert 1997, pp. 894-896). These facts strongly indicated that the hammer and anvil had evolved from these reptilian jawbones—that is, if common descent was in fact true. This result was so striking, and the required intermediates so outlandish, that many anatomists had extreme trouble imagining how transitional forms bridging these morphologies could have existed while retaining function. Young-earth creationist Duane Gish stated the problem this way:
|
"All mammals, living or fossil, have a single bone, the dentary, on each side of the lower jaw, and all mammals, living or fossil, have three auditory ossicles or ear bones, the malleus, incus and stapes. ... Every reptile, living or fossil, however, has at least four bones in the lower jaw and only one auditory ossicle, the stapes. ... There are no transitional fossil forms showing, for instance, three or two jawbones, or two ear bones. No one has explained yet, for that matter, how the transitional form would have managed to chew while his jaw was being unhinged and rearticulated, or how he would hear while dragging two of his jaw bones up into his ear." (Gish 1978, p. 80) |
![[Figure1.4.2a (cartoon of vertebrate ears)]](http://www.talkorigins.org/faqs/comdesc/images/reptile_ear.gif) ![[Figure1.4.2b (cartoon of vertebrate ears)]](http://www.talkorigins.org/faqs/comdesc/images/human_ear.gif) |
|
Figure 1.4.2. A comparison of the ears of reptiles and mammals. The reptile ear is shown on the left, the mammal ear on the right. As in Figure 1.4.1, the quadrate (mammalian anvil or incus) is in turquoise and the articular (mammalian hammer or malleus) is in yellow. The stapes is shown in brown. Note how the relative arrangement of these bones is similar in both taxa, in the order of inner ear-stapes-quadrate-articular. |
Gish was incorrect in stating that there were no transitional fossil forms, and he has been corrected on this gaff numerous times since he wrote these words. However, Gish's statements nicely delineate the morphological conundrum at hand. Let's review the required evolutionary conclusion. During their evolution, two mammalian middle ear bones (the hammer and anvil, aka malleus and incus) were derived from two reptilian jawbones. Thus there was a major evolutionary transition in which several reptilian jawbones (the quadrate, articular, and angular) were extensively reduced and modified gradually to form the modern mammalian middle ear. At the same time, the dentary bone, a part of the reptilian jaw, was expanded to form the major mammalian lower jawbone. During the course of this change, the bones that form the hinge joint of the jaw changed identity. Importantly, the reptilian jaw joint is formed at the intersection of the quadrate and articular whereas the mammalian jaw joint is formed at the intersection of the squamosal and dentary (see Figure 1.4.1).
How could hearing and jaw articulation be preserved during this transition? As clearly shown from the many transitional fossils that have been found (see Figure 1.4.3), the bones that transfer sound in the reptilian and mammalian ear were in contact with each other throughout the evolution of this transition. In reptiles, the stapes contacts the quadrate, which in turn contacts the articular. In mammals, the stapes contacts the incus, which in turn contacts the malleus (see Figure 1.4.2). Since the quadrate evolved into the incus, and the articular evolved into the malleus, these three bones were in constant contact during this impressive evolutionary change. Furthermore, a functional jaw joint was maintained by redundancy—several of the intermediate fossils have both a reptilian jaw joint (from the quadrate and articular) and a mammalian jaw joint (from the dentary and squamosal). Several late cynodonts and Morganucodon clearly have a double-jointed jaw. In this way, the reptilian-style jaw joint was freed to evolve a new specialized function in the middle ear. It is worthy of note that some modern species of snakes have a double-jointed jaw involving different bones, so such a mechanical arrangement is certainly possible and functional.
Since Figure 1.4.3 was made, several important intermediate fossils have been discovered that fit between Morganucodon and the earliest mammals. These new discoveries include a complete skull of Hadrocodium wui (Luo et al. 2001) and cranial and jaw material from Repenomamus and Gobiconodon (Wang et al. 2001). These new fossil finds clarify exactly when and how the malleus, incus, and angular completely detached from the lower jaw and became solely auditory ear ossicles.
Recall that Gish stated: "There are no transitional fossil forms showing, for instance, three or two jawbones, or two ear bones" (Gish 1978, p. 80). Gish simply does not understand how gradual transitions happen (something he should understand, obviously, if he intends to criticize evolutionary theory). These fossil intermediates illustrate why Gish's statement is a gross mischaracterization of how a transitional form should look. In several of the known intermediates, the bones have overlapping functions, and one bone can be called both an ear bone and a jaw bone; these bones serve two functions. Thus, there is no reason to expect transitional forms with intermediate numbers of jaw bones or ear bones. For example, in Morganucodon, the quadrate (anvil) and the articular (hammer) serve as mammalian-style ear bones and reptilian jaw bones simultaneously. In fact, even in modern reptiles the quadrate and articular serve to transmit sound to the stapes and the inner ear (see Figure 1.4.2). The relevant transition, then, is a process where the ear bones, initially located in the lower jaw, become specialized in function by eventually detaching from the lower jaw and moving closer to the inner ear.
![[Figure1.4.3 (cartoon of vertebrate jaws)]](http://www.talkorigins.org/faqs/comdesc/images/jaws1.gif) Figure 1.4.3. A comparison of the jawbones and ear-bones of several transitional forms in the evolution of mammals. Approximate stratigraphic ranges of the various taxa are indicated at the far left (more recent on top). The left column of jawbones shows the view of the left jawbone from the inside of the mouth. The right column is the view of the right jawbone from the right side (outside of the skull). As in Figure 1.4.1, the quadrate (mammalian anvil or incus) is in turquoise, the articular (mammalian hammer or malleus) is in yellow, and the angular (mammalian tympanic annulus) is in pink. For clarity, the teeth are not shown, and the squamosal upper jawbone is omitted (it replaces the quadrate in the mammalian jaw joint, and forms part of the jaw joint in advanced cynodonts and Morganucodon). Q = quadrate, Ar = articular, An = angular, I = incus (anvil), Ma = malleus (hammer), Ty = tympanic annulus, D = dentary. (Reproduced from Kardong 2002, pp. 274, with permission from the publisher, Copyright © 2002 McGraw-Hill) |
The above is from
Mammal-Like Reptiles
As previously stated, a succession of transitional fossils exists that link reptiles (Class Reptilia) and mammals (Class Mammalia). These particular reptiles are classifie as Subclass Synapsida. Presently, this is the best example of th e transformation of one major higher taxon into another. The morphologic changes that took place are well documented by fossils, beginning with animals essentially 100% reptilian and resulting in animals essentially 100% mammalian. Therefore, I have chosen this as the example to summarize in more detail (Table 1, Fig. 1).
![[Fig. 1a]](http://www.gcssepm.org/images/fossil_a.gif)
![[Fig. 1b]](http://www.gcssepm.org/images/fossil_b.gif)
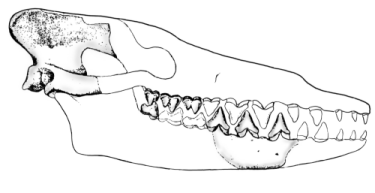
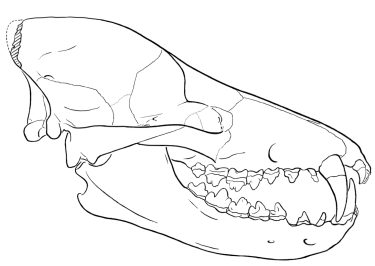
Skulls and jaws of synapsid reptiles and mammals; left column side view of skull; center column top view of skull; right column side view of lower jaw. Hylonomus modified from Carroll (1964, Figs. 2,6; 1968, Figs. 10-2, 10-5; note that Hylonomus is a protorothyrod, not a synapsid). Archaeothyris modified from Reisz (1972, Fig. 2). Haptodus modified from Currie (1977, Figs, 1a, 1b; 1979, Figs. 5a, 5b). Sphenacodo n modified from Romer & Price (1940, Fig. 4f), Allin (1975, p. 3, Fig. 16);note: Dimetrodon substituted for top view; modified from Romer & Price, 1940, pl. 10. Biarmosuchus modified from Ivakhnenko et al. (1997, pl. 65, Figs. 1a, 1B, 2); Alin & Hopson (1992; Fig. 28.4c); Sigogneau & Tchudinov (1972, Figs. 1, 15). Eoarctops modified from Broom (1932, Fig. 35a); Boonstra (1969, Fig. 18). Pristerognathus modified from Broom (1932, Figs 17a, b,c); Boonstra (1963, Fig. 5d). Procynosuchus modified from Allin & Hopson (1992, Fig. 28.4e); Hopson (1987, Fig. 5c); Brink (1963, Fig. 10a); Kemp (1979, Fig. 1); Allin (1975, p. 3, Fig. 14). Thrinaxodon modified from Allin & Hopson (1992, Fig. 28.4f);Parrington (1946, Fig. 1); Allin (1975, p. 3, Fig. 13). Probainognathus modified from Allin & Hopson (1992, Fig. 28.4g); Romer (1970, Fig. 1); Allin (1975, p. 3, Fig. 12). Morga nucodon modified from Kermack, Mussett, & Rigney (1981, Figs. 95, 99a; 1973, Fig. 7a); Allin (1975, p. 3, Fig. 11). Asioryctes modified from Carroll (1988, Fig. 20-3b). Abbreviations: ag = angular; ar = articular; cp = coronoid process; d = dentary; f = lateral temporal fenestra; j = jugal; mm = attachment site for mammalian jaw muscles; o = eye socket; po = post orbital; q = quadrate; rl = reflected lamina; sq = squamosal; ty = tympanic.
|
TAXONOMY
|
LATERAL TEMPORAL FENESTRA
|
LOWER JAW DENTARY
|
TEETH
|
LOWER JAW: POST- DENTARY BONES
|
MIDDLE EAR & JAW ARTICULATION
|
M: Early Placental mammals
Asioryctes
Upper Cretaceous |
Merged with eye socket; cheek arch bowed out laterally |
100% of jaw length is the den- tary; condylar process in contact with squamosal |
Fully differentiated teeth; incisors, canines, premolars; one tooth replacement |
No post-dentary bones |
3 middle ear bones (stapes, incus, malleus) + tympanic; squamosal-dentary jaw joint |
L: "Pantothere" mammals
Amphitherium
Middle/Upper Jurassic |
X |
100% of jaw length is the den- tary; condylar process contacts squamosal |
Fully differentiated teeth; incisors, canines, premolars; one tooth replacement |
Post-dentary bones migrated to middle ear |
Probably 3 middle ear bones (stapes, incus, malleus) + tympanic; squamosal-dentary jaw joint |
K: Morganucodontid mammals
Morganucodon Upper Triassic & Lower Jurassic |
Merged with eye socket; cheeck arch bowed out laterally |
100% of jaw length is the den- tary; condylar process expanded posteriorly to make contact with squamosal |
Fully differentiated teeth; incisors, canines, premolars; one tooth replacement |
20% of jaw length; reflected lamina decreased to narrow ribbon-like horseshoe |
Stapes extends from inner ear capsule to quadrate; quadrate tiny; both quadrate-articular and squamosal-dentary jaw joints |
J: Chiniquodontid cynodonts
Probainognathus
Middle Triassic |
Much larger than eye socket; 40- 45% of skull length; expanded posterioirly, medially, & laterally; midline of skull narrow sagittal crest; chek arch bowed out laterally |
95% of jaw length is the dentary; large coronoid process expanded posteriorly; condylar process expanded posteriorly |
Large single canine; cheek teeth multicusped; tooth replacement reduced |
20% of jaw length; angular notch widened ventrally; width of main part of angular decreased; reflec - ted lamina decreased to narrow ribbon-like horseshoe |
Stapes extends from inner ear capsule to quadrate; quadrate tiny; quadrate-articular joint |
I:Galesaurid cynodonts
Thrinaxodon
Lower Triassic |
Much larger than eye socket; 40% of skull length; expanded pos- terioirly, medially, & laterally; midline of skull narrow sagittal crest; chek arch bowed out laterally |
85% of jaw length is the dentary; large coronoid process expanded to top of eye socket and pos- teriorly; jaw muscles attached to most of coronoid process |
Large single canine; cheek teeth multicusped; tooth replacement reduced |
25% of jaw length; angular notch widened ventrally; width of reflec- ted lamina decreased; width of main part of angular decreased |
Stapes extends from inner ear capsule to quadrate; quadrate small; quadrate-articular jaw joint |
H: Procynosuchid cynodonts
Procynosuchus
upper Upper Permian |
Much larger than eye socket; 40% of skull length; expanded pos- terioirly, medially, & laterally; midline of skull narrow sagittal crest; chek arch bowed out laterally |
75-80% of jaw length is the den- tary; coronoid process expanded to near top of eye socket and posteriorly; jaw muscles attached to dorsal part of coronoid process |
Large single canine; cheek teeth multicusped |
30% of jaw length; angular notch widened ventrally; width of reflected lamina decreased |
Stapes extends from inner ear capsule to quadrate; quadrate small; quadrate-articular jaw joint |
G: Early Therocephalians
Pristerognathus
lower Upper Permian |
Larger than eye socket; expanded posteriorly and medially; 30% of skull length |
75-80% of jaw length is the den- tary; posterior end of dentary expanded posteriorly and dorsally into narrow blade-like coronoid process; rises to middle of eye socket |
Large single canine; other teeth simple cones. |
35% of jaw length; angular notch deepened into a cleft; reflected lamina large, broad, blade-like |
Stapes extends from inner ear capsule to quadrate; quadrate small; quadrate-articular jaw joint |
F: Early Gorgonopsians
Eoarctops
lower Upper Permian |
Slightly larger than eye socket; expanded posteriorly and medially (minimal); 20-25% of skull length |
65-75% of jaw length is the den- tary; posterior end of dentary slightly expanded posteriorly and dorsally as incipient coronoid process |
Large single canine; other teeth simple cones. |
40% of jaw length; angular notch deepened into a cleft; reflected lamina large, broad, blade-like |
Stapes extends from inner ear capsule to quadrate; quadrate- articular jaw joint |
E: Eotitanosuchians
Sphenacodon
Lower Permian |
Small; slightly smaller than eye socket; slightly expanded posteriorly and medially |
65-75% of jaw length is the den- tary; posterodorsal edge rises broadly but slightly above tooth row |
Large single canine; other teeth simple cones. |
40% of jaw length; angular notch deepened into a cleft; reflected lamina large, broad, blade-like |
Stapes extends from inner ear capsule to quadrate; quadrate- articular jaw joint |
D: Late sphenacodonts
Sphenacodon
Upper Pennsylvanian |
Small; smaller than eye socket; confined to one side of skull |
65% of jaw length is the dentary; posterodorsal edge rises broadly but slightly above the tooth row |
Enlarged incipient canines; other teeth simple cones |
60% of jaw length; venntral edge of angular notched ("angular notch") offsetting a short pro- tusion (reflected lamina) |
Stapes extends from inner ear capsule to quadrate; quadrate large and plate-like; quadrate- articular jaw joint |
C: Early spenacodonts
Haptodus
Upper Pennsylvanian |
Tiny; smaller than eye socket; confined to one side of skull |
65-75% of jaw length is the den- tary; posterodorsal edge rises broadly but slightly above tooth row |
Undifferentiated; slightly enlarged incipient canines just behind nares |
70% of jaw length; ventral edge of angular with shallow indentation |
Stapes extends from inner ear capsule to quadrate; quadrate- articular jaw joint |
B: Early ophiacodonts
Archaothyris
upper Middle Pennsylvanian |
Tiny; smaller than eye socket; confined to one side of skull |
x |
Undifferentiated; slightly enlarged incipient canines just behind nares |
x |
Stapes extends from inner ear capsule to quadrate; quadrate- articular jaw joint |
A: Protorothyrids
Hylonomus
lower Middle Pennsylvanian |
Absent |
65-75% of jaw length is the den- tary; posterodorsal edge rises broadly but slightly above tooth row |
Undifferentiated; slightly enlarged incipient canines just behind nares |
70% of jaw length; ventral edge of angular continuous |
Stapes extends from inner ear capsule to quadrate; quadrate- articular jaw joint |
Table 1: Morphology of synapsid reptiles and mammals (Note that Hylonomus is a protothyrid, not a synapsid). Data from references cited in text.
Modern reptiles and mammals are very distinctive, easily diagnosable, and do not intergrade. Reptiles are covered by scales, mammals by hair; reptiles are cold-blooded, mammals warm-blooded; reptiles do not suckle their young, mammals have mammary glands; reptiles have sprawling posture, mammals have upright posture. Most of these features are soft part anatomy or physiology that very rarely fossilize (although dinosaur skin impressions are known from Cretaceous sediments, and imprints of mammal hair are known from Eocene bats from Germany; Franzen, 1990). In the fossil record, we must look to skeletal features.
There are many skeletal features which allow us to distinguish the reptiles from the mammals (Carroll, 1988; Table 1, rows A, M). The single most important defining characteristic is the nature of the articulation of the lower jaw to the skull (Simpson, 1959). In reptiles, multiple bones comprise the lower jaw. A small bone at the posterior end of the lower jaw, the articular, articulates with the quadrate bone of the skull (Simpson, 1959; Carroll, 1988). In mammals, one large bone, the dentary, comprises the lower jaw. It articulates with the squamosal bone of the skull (Simpson, 1959; Carroll, 1988).
From comparative anatomy studies, it is certain that most of the bones of the reptiles and mammals are homologous (Crompton & Parker, 1978; Carroll, 1988). Of greatest importance, the middle ear bones of mammals (stapes, incus, malleus, and tympanic) are homologous with several of the skull and jaw bones of reptiles (stapes, quadrate, articular, and angular, respectively; Romer, 1956, p. 33-38, 1970a; Allin, 1975, 1986; Allin & Hopson, 1992; Crompton & Parker, 1978; Hopso n, 1987, 1994; Carroll, 1988). One group of reptiles, the synapsids (Subclass Synapsida), share with the mammals an additional homologous structure: the lateral temporal fenestra, which is an opening in the skull behind the eye socket at the triple junction between the squamosal, jugal , and post orbital bones (Broom, 1932; Frazetta, 1968; Kemp, 1982; Carroll, 1988). A band of bone composed of the jugal and the squamosal is adjacent to the lateral temporal fenestra (Broom, 1932; Kemp, 1982; Carroll, 1988). This is the cheek arch so characteristic of mammal skulls (Broom, 1932; Kemp, 1982; Carroll, 1988). Therefore, synapsids are commonly named the “mammal-like reptiles.”
The presence of diagnosable morphologic differences between reptiles (including the oldest reptiles and the oldest synapsids) and mammals distinguishes them as distinct taxa. This allows us to test evolution by looking for transitional forms between the two. Because many of the bones are homologous, we should find evidence illustrating how these bones were modified over time to become the new bones. Furthermore, these morphologic changes should happen in parallel and in geochronologic succession.
Synapsid reptiles inhabited Pangea from the Middle Pennsylvanian through the Early Jurassic (Kemp, 1982, 1985; Sloan, 1983; Carroll, 1988; Hopson, 1969, 1987, 1994; Hopson & Crompton, 1969; Hotton, et al., 1986; Crompton & Jenkins, 1973; Sidor & Hopson, 1998; Romer & Price, 1940; Broom, 1932; Boonstra, 1963, 1969, 1971; Tchudinov, 1983; Olson, 1944; Tatarinov, 1974; Vyushkov, 1955; Efremov, 1954). From the Early Permian through the Early Triassic, they were the largest and most abundant land animals (Sloan, 1983; Colbert, 1965). Though much less well known to the general public than dinosaurs, one of the “cereal box dinosaurs,” Dimetrodon (the sail-backed reptile), is a synapsid, not a dinosaur (Romer & Price, 1940; Carroll, 1988). The oldest mammals are Late Triassic (Kemp, 1982; Carroll, 1988). Below is a discussion of the geochronologic succession linking synapsids and mammals. The oldest reptiles (named protorothyrids; Carroll, 1964, 1988, p. 192-199) are from the lower Middle Pennsylvanian, and the oldest synapsids (Reisz, 1972) are from the upper Middle Pennsylvanian, both of Nova Scotia. Upper Pennsylvanian and Lower Permian forms are known primarily from the midcontinent and Permian Basin region of the United States (Romer & Price, 1940; Currie, 1977, 1979; Kemp, 1982; Sloan, 1983). The basal Upper Permian forms are known from Russia (Tchudinov, 1960, 1983; Efremov, 1954; Olson, 1962; Sigogneau & Tchudinov, 1972; Ivakhnenko et al., 1997). Most of the Upper Permian and Lower Triassic succession is known from southern Africa, especially the Great Karoo of South Africa (Broom, 1932; Boonstra, 1963, 1969, 1971; Hopson & Kitching, 1972; Kemp, 1982; Sloan, 1983). The Middle Triassic forms are from South America (Romer, 1969a, 1969b, 1970b, 1973; Romer & Lewis, 1973; Bonaparte & Barbarena, 1975), and the Upper Triassic and Lower Jurassic mammals are known from Eurasia (Kermack, Mussett, & Rigney, 1973, 1981; Kemp, 1982). Subsequent Mesozoic mammals are known from all over the world (Simpson, 1928; Lillegraven et al., 1979).
When placed in proper geochronologic succession, the synapsids naturally form a succession of taxa (genera and families) that progressively become more mammal-like and less reptile-like (Kemp, 1982, 1985; Sloan, 1983; Sidor & Hopson, 1998; Hopson, 1987, 1994). Morphologic changes, summarized in Table 1 and Figure 1, affect the entire skeletal anatomy of these animals, but are most clearly displayed in their skulls.
The lateral temporal fenestra increased in size from a tiny opening smaller than the eye socket to a giant opening occupying nearly half the length of the skull. Ultimately, it merged with the eye socket, thus producing the full development of the cheek arch so characteristic of mammals (Broom, 1932; Frazetta, 1968; Kemp, 1982; Sloan, 1983; Hopson, 1987, 1994; Carroll, 1988).
Successively, the relative proportion of the lower jaw comprised of the dentary bone (teeth-bearing bone) gradually increased until the entire lower jaw consisted of the dentary (Kemp, 1982; Sloan, 1983; Carroll, 1988; Hopson, 1987, 1994). In Pennsylvanian and Lower and basal Upper Permian synapsids, the postero-dorsal edge of the lower jaw rose broadly but only slightly above the level of the tooth row (Romer & Price, 1940; Currie, 1977, 1979; Ivakhnenko et al., 1997; Tchudinov, 1960, 1983; Efremov, 1954; Olson, 1962; Sigogneau & Tchudinov, 1972; Hopson, 1987, 1994). In succeeding forms, the posterior part of the dentary expanded dorsally and posteriorly as a blade-like process, and progressively became larger (Broom, 1932; Boonstra, 1963, 1969, 1971; Sigogneau, 1970; Brink, 1963; Kemp, 1979; Hopson, 1987, 1994), forming the coronoid process (Parrington, 1946; Fourie, 1974; Romer, 1969b, 1970b, 1973; Hopson, 1987, 1994) to which the mammalian-type jaw musculature is attached (Barghusen, 1968; Bramble, 1978; Crompton, 1972; Crompton & Parker, 1978; Kemp, 1982; Sloan, 1983; Carroll, 1988). Concomitantly, the post-dentary bones progressively reduced in size (Allin, 1975; Crompton, 1972; Crompton & Parker, 1978; Kemp, 1982; Sloan, 1983; Carroll, 1988; Hopson, 1987, 1994).
Beginning with the Upper Pennsylvanian sphenacodonts, a notch developed in the angular bone that offsets a projection, the reflected lamina (Allin, 1975; Allin & Hopson, 1992; Hopson, 1987, 1994; Romer & Price, 1940; Currie, 1977, 1979; Kemp, 1982; Sloan, 1983; Carroll, 1988). The reflected lamina first became a large blade-like flange (Allin, 1975; Allin & Hopson, 1992; Hopson, 1987, 1994; Ivakhnenko et al., 1997; Tchudinov, 1960, 1983; Efremov, 1954; Olson, 1962; Sigogneau & Tchudinov, 1972; Broom, 1932; Sigogneau, 1970; Boonstra, 1963, 1969, 1971), and then was progressively reduced to a delicate horseshoe-shaped bone (Allin, 1975; Allin & Hopson, 1992; Hopson, 1987, 1994; Brink, 1963; Parrington, 1946; Fourie, 1974; Romer, 1969b, 1970b, 1973; Kermack, Mussett, & Rigney, 1973, 1981; Kemp, 1979, 1982; Sloan, 1983; Carroll, 1988).
Simultaneously, the quadrate progressively decreased in size (Allin, 1975; Allin & Hopson, 1992; Hopson, 1987, 1994; Kemp, 1982; Sloan, 1983; Carroll, 1988). The articular did not decrease in size much, being small initially, but developed a downward-pointing prong (Allin, 1975; Allin & Hopson, 1992; Hopson, 1987, 1994; Kemp, 1982; Sloan, 1983; Carroll, 1988). In the synapsids, the lower jaw was hinged to the skull by the articular and quadrate bones (Crompton, 1972; Crompton & Parker, 1978; Allin, 1975; Allin & Hopson, 1992; Hopson, 1987, 1994). Thus they are classified as reptiles (Simpson, 1959; Kemp, 1982; Sloan, 1983; Carroll, 1988). As the quadrate and articular became smaller, they were relieved of their solid suture to the dentary and skull (Crompton, 1972; Allin, 1975, 1986; Allin & Hopson, 1992; Hopson, 1987, 1994; Crompton & Parker, 1978; Kemp, 1982; Sloan, 1983; Carroll, 1988). A projection of the dentary extended posteriorly and made contact with the squamosal. Morganucodon possessed the mammalian dentary-squamosal jaw joint adjacent to the reptilian articular-quadrate jaw joint (Kermack, Mussett, & Rigney, 1973, 1981; Carroll, 1988). It is classified as the first mammal, but it is a perfect intermediate. Now that a new jaw joint was established, the quadrate and articular were subsequently relieved of that function (Crompton, 1972; Allin, 1975, 1986; Allin & Hopson, 1992; Hopson, 1987, 1994; Crompton & Parker, 1978; Kemp, 1982; Sloan, 1983; Carroll, 1988). Ultimately, in Middle and Upper Jurassic mammals, the tiny quadrate, articular, and ring-like angular migrated as a unit to the middle ear where they joined the stapes and became the incus, malleus, and tympanic bones (Allin, 197 5, 1986; Allin & Hopson, 1992; Hopson, 1987, 1994; Kemp, 1982; Sloan, 1983; Carroll, 1988).
Progressively, the teeth became differentiated. The large canines developed first, followed by the development of multicusped cheek teeth, reduced tooth replacement (Osborn & Crompton, 1973; Crompton & Parker, 1978), and finally full y differentiated incisors, canines, premolars, and molars with one tooth replacement during life (Kemp, 1982; Hopson, 1994).
Many other morphologic changes are documented in the fossil record. These demonstrate the morphologic and geochronologic succession from sprawling limb posture to upright limb posture of mammals (Jenkins, 1971; Romer & Lewis, 197 3; Kemp, 1982; Carroll, 1988; Hopson, 1994). As Jenkins (1971, p. 210) stated, “In details of morphology and function, the cynodont post-cranial skeleton should be regarded as neither ‘reptilian’ nor ‘mammalian’ but as transitional between the two classes .” Other changes have been adequately summarized elsewhere (Kemp, 1982; Sloan, 1983; Carroll, 1988; Hopson, 1994). Obviously, fundamental physiologic changes must have taken place as well, many of which are not directly preserved in the fossil record, though some can be inferred from the skeletal anatomy (Findlay, 1968; Kemp, 1982; Sloan, 1983, Carroll, 1988; Hopson, 1994).
This is well documented in the fossil record by a massive volume of incontrovertible data that cannot be explained away. Such large-scale, progressive, continuous, gradual, and geochronologically successive morphologic change (Sidor & Hopson, 1998) is descent with modification, and provides compelling evidence for evolution on a grand scale.
(The above is from volution, the overarching concept that unifies the biological sciences, in fact embraces a plurality of theories and hypotheses. In evolutionary debates one is apt to hear evolution roughly parceled between the terms "microevolution" and "macroevolution". Microevolution, or change beneath the species level, may be thought of as relatively small scale change in the functional and genetic constituencies of populations of organisms. That this occurs and has been observed is generally undisputed by critics of evolution. What is vigorously challenged, however, is macroevolution. Macroevolution is evolution on the "grand scale" resulting in the origin of higher taxa. In evolutionary theory it thus entails common ancestry, descent with modification, speciation, the genealogical relatedness of all life, transformation of species, and large scale functional and structural changes of populations through time, all at or above the species level (Freeman and Herron 2004; Futuyma 1998; Ridley 1993).




















![[Figure1.4.1 (cartoon of vertebrate jaws)]](http://www.talkorigins.org/faqs/comdesc/images/jaws2.gif)
![[Figure1.4.2a (cartoon of vertebrate ears)]](http://www.talkorigins.org/faqs/comdesc/images/reptile_ear.gif)
![[Figure1.4.2b (cartoon of vertebrate ears)]](http://www.talkorigins.org/faqs/comdesc/images/human_ear.gif)
![[Figure1.4.3 (cartoon of vertebrate jaws)]](http://www.talkorigins.org/faqs/comdesc/images/jaws1.gif)
![[Fig. 1a]](http://www.gcssepm.org/images/fossil_a.gif)
![[Fig. 1b]](http://www.gcssepm.org/images/fossil_b.gif)